Modeling Non-linear relationships
Modeling Non-linear relationships
Learning objectives:
Be able to implement common approaches for modeling non-linear relationships between \(X_i\) and \(Y_i\)
- Polynomials using the
polyfunction in R - Splines using the
nsfunction (splines library) - Smoothing splines
- Polynomials using the
- Understand how model predictions are constructed when using polynomials or splines
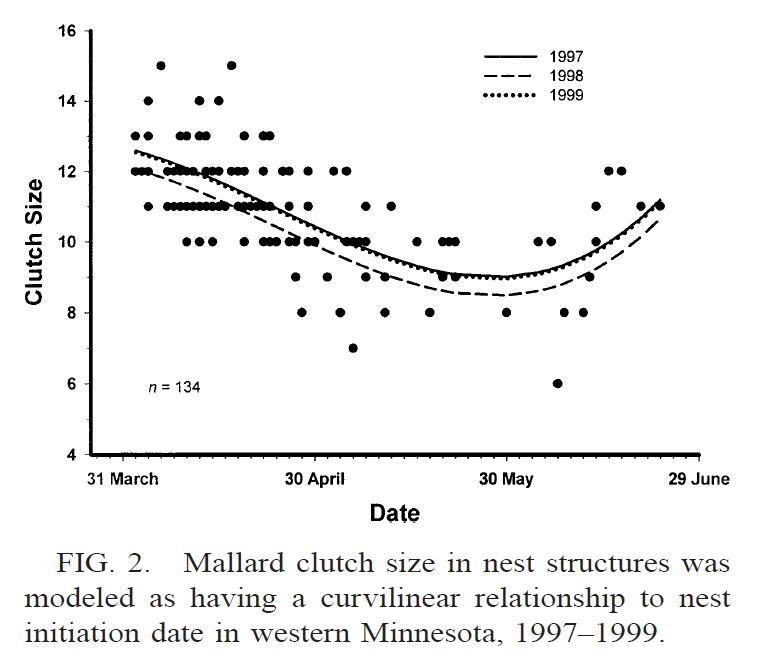
Mallard clutch size versus Julian Date

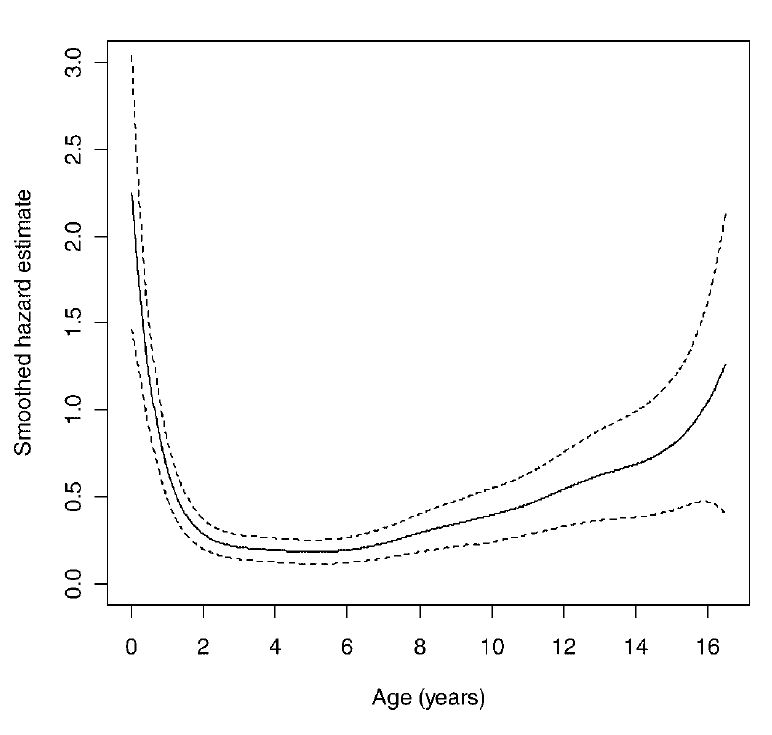
Age-specific Hazard for White-tailed Deer

Linear Models
So far, we have focused on linear models of the form:
\(Y_i = \beta_0 + X_i\beta + \epsilon_i\)
or
\(Y_i = \beta_0 + X_{i,1}\beta_1 + X_{i,2}\beta_2 + \ldots + \epsilon_i\)
The model can be written as a “linear combination” of parameters.
Modeling Non-Linear Relationships
- Polynomials (e.g., poly(age,2) for a quadratic in age)
- Transformations of \(X\) or \(Y\) (e.g., log(\(X\)), \(\sqrt{Y}\), \(exp(X)\)).
- Regression splines
These options still lead to linear models:
\(Y_i = \beta_0 + X_i\beta_1 + X^2_i\beta_2+ \ldots + \epsilon_i\)
\(\sqrt{Y_i} = \beta_0 + log(X_i)\beta_1 + \ldots + \epsilon_i\)
So, we can use all the same tools we’ve learned about (e.g., residual plots, t-tests, F-tests, AIC, etc) [note: try writing out the above models in matrix notation!]
Species-area relationship
Plant species richness for 29 islands in the Galapagos Islands archipelago (Johnson and Raven 1973)
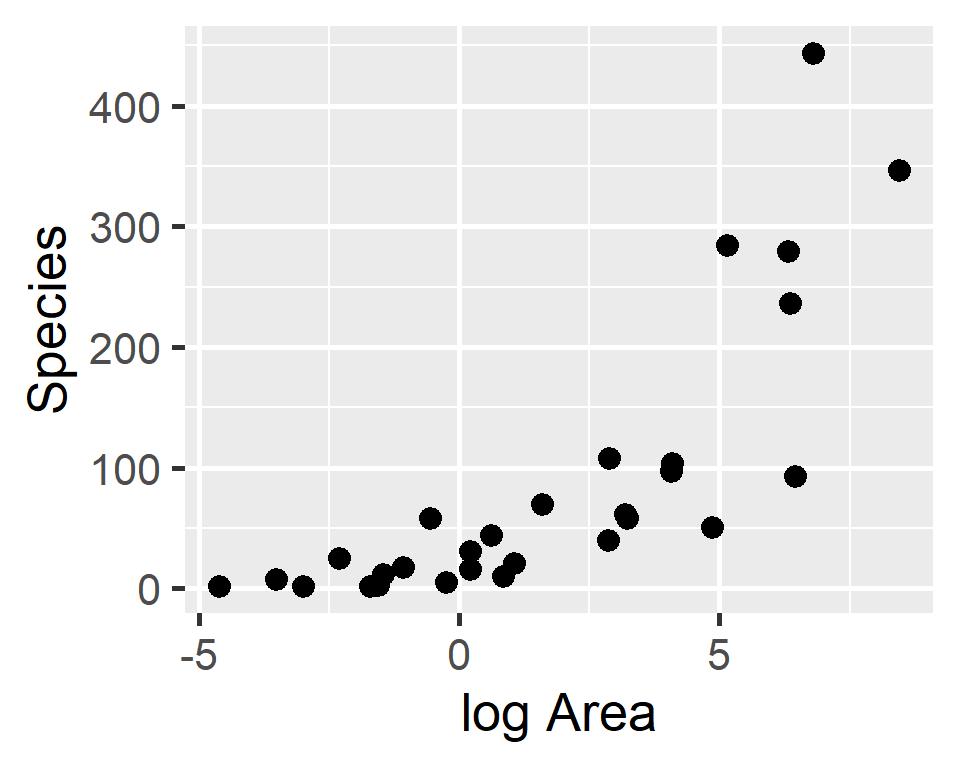
Species-area relationship
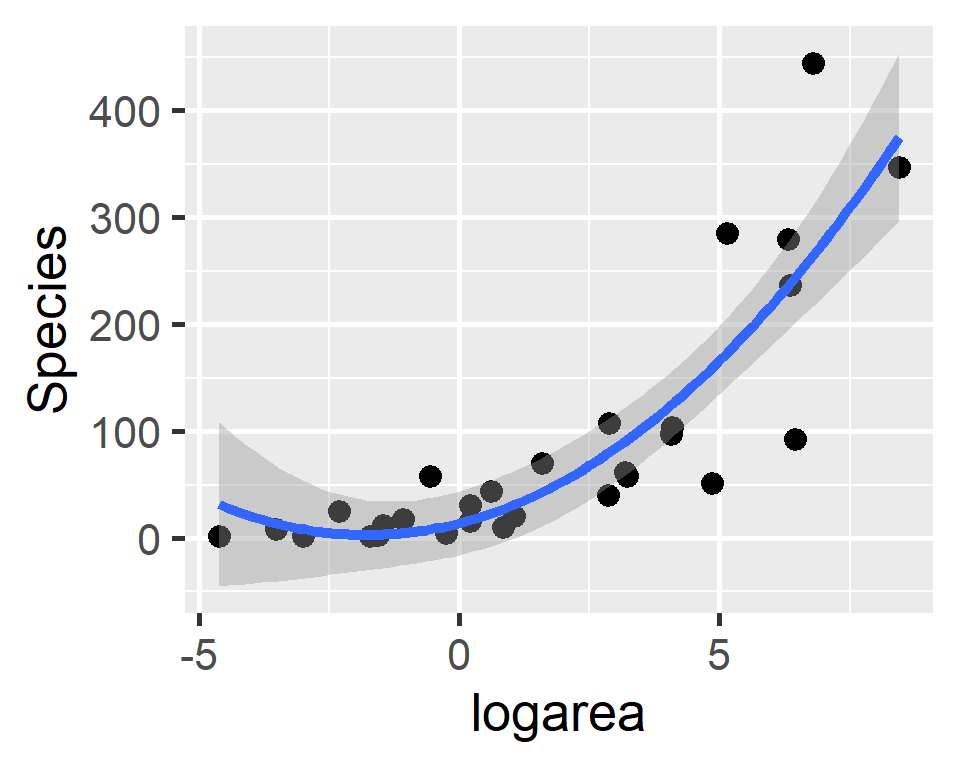
Polynomials
Call:
lm(formula = Species ~ logarea + logarea.squared, data = gala)
Residuals:
Min 1Q Median 3Q Max
-151.009 -27.361 -1.033 20.825 178.805
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 14.1530 14.5607 0.972 0.340010
logarea 12.6226 4.8614 2.596 0.015293 *
logarea.squared 3.5641 0.9445 3.773 0.000842 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 59.88 on 26 degrees of freedom
Multiple R-squared: 0.7528, Adjusted R-squared: 0.7338
F-statistic: 39.6 on 2 and 26 DF, p-value: 1.285e-08Polynomials
Call:
lm(formula = Species ~ poly(logarea, 2, raw = TRUE), data = gala)
Residuals:
Min 1Q Median 3Q Max
-151.009 -27.361 -1.033 20.825 178.805
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 14.1530 14.5607 0.972 0.340010
poly(logarea, 2, raw = TRUE)1 12.6226 4.8614 2.596 0.015293 *
poly(logarea, 2, raw = TRUE)2 3.5641 0.9445 3.773 0.000842 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 59.88 on 26 degrees of freedom
Multiple R-squared: 0.7528, Adjusted R-squared: 0.7338
F-statistic: 39.6 on 2 and 26 DF, p-value: 1.285e-08Polynomials: component + residual plot
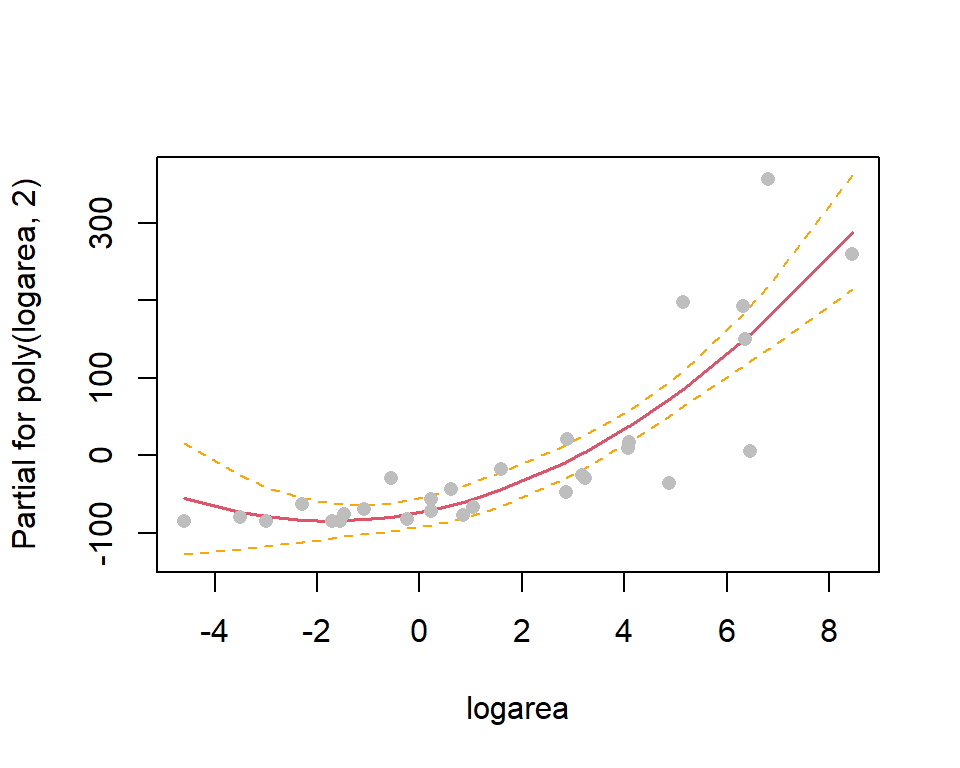
Hypothesis Testing
Anova Table (Type II tests)
Response: Species
Sum Sq Df F value Pr(>F)
logarea 24175 1 6.7417 0.0152925 *
logarea.squared 51058 1 14.2387 0.0008418 ***
Residuals 93232 26
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Basis functions/vectors
A linear model is a model that is linear in the parameters:
\(Y_i = \sum_{j=1}^P \beta_j b_j(X_i) + \epsilon_i\)
The \(b_j(X_i)\) are called basis functions or basis vectors.
\(Y_i = \beta_0 + \beta_2X_i + \beta_3X_i^2 + \ldots + \epsilon_i\)
\(b_j(X_i) = 1, X, X^2, X^3, \ldots\)
Basis functions
\(E[Y_i|X_i] = \beta_01 + \beta_2X_i + \beta_3X_i^2\)
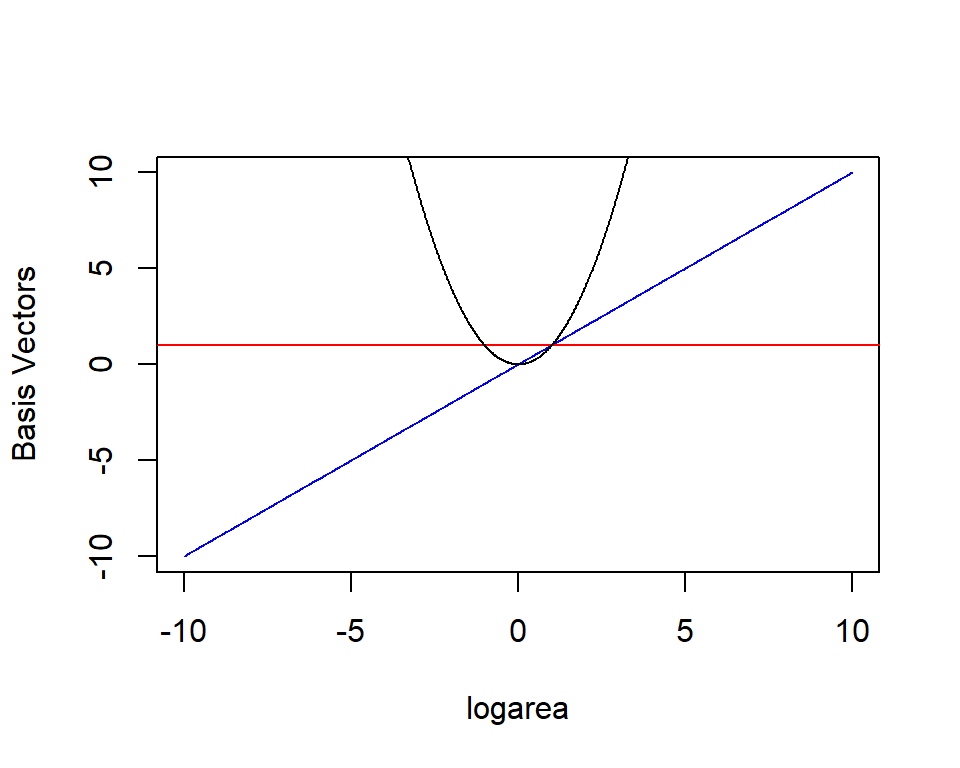
\(E[Y | X]\) is given by a linear combination of a horizontal line (1), a line through the origin (\(X\)), a quadratic centered on the origin (\(X^2\)).
Species-Area relationship
\(Species_i = 14.15 + 12.62X_i + 3.56X^2_i\)
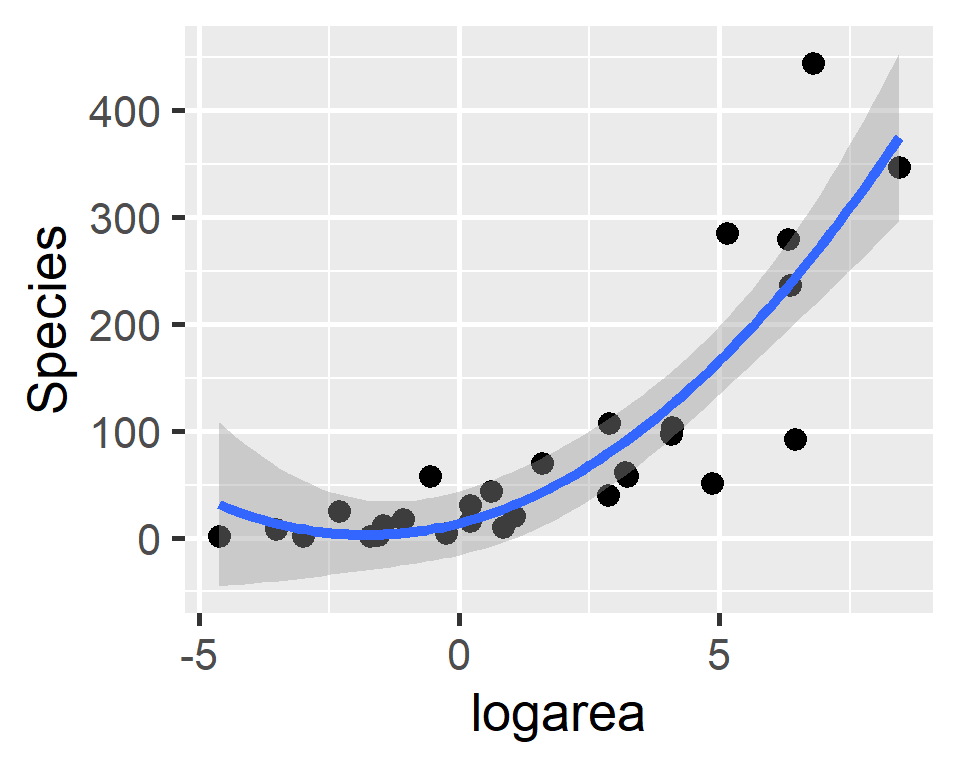
Polynomials
A polynomial of degree D is a function formed by linear combinations of the powers of its argument up to D:
\(y = \beta_0 + \beta_1x + \beta_2x^2 + \ldots + \beta_Dx^D\)
Specific polynomials:
- Linear: \(y = \beta_0 + \beta_1x\)
- Quadratic:: \(y = \beta_0 + \beta_1x + \beta_2x^2\)
- Cubic: \(y = \beta_0 + \beta_1x + \beta_2x^2 + \beta_3x^3\)
- Quartic: \(y = \beta_0 + \beta_1x + \beta_2x^2 + \beta_3x^3+\beta_4x^4\)
- Quintic: \(y = \beta_0 + \beta_1x + \beta_2x^2 + \beta_3x^3+\beta_4x^4 + \beta_5x^5\)
Polynomials
The design matrix for a regression model with \(n\) observations and \(p\) predictors is the matrix with \(n\) rows and \(p\) columns such that the value of the \(j^{th}\) predictor for the \(i^{th}\) observation is located in column \(j\) of row \(i\).
Design matrix for a polynomial of degree D
\(\begin{bmatrix} 1 & x_1 & x_1^2 & x_1^3 & ... & x_1^D \\ 1 & x_2 & x_2^2 & x_2^3 & ... & x_2^D \\ 1 & x_3 & x_3^2 & x_3^3 & ... & x_3^D \\ & & \vdots & & \\ 1 & x_n & x_n^2 & x_n^3 & ... & x_n^D \\ \end{bmatrix}\)
Polynomials
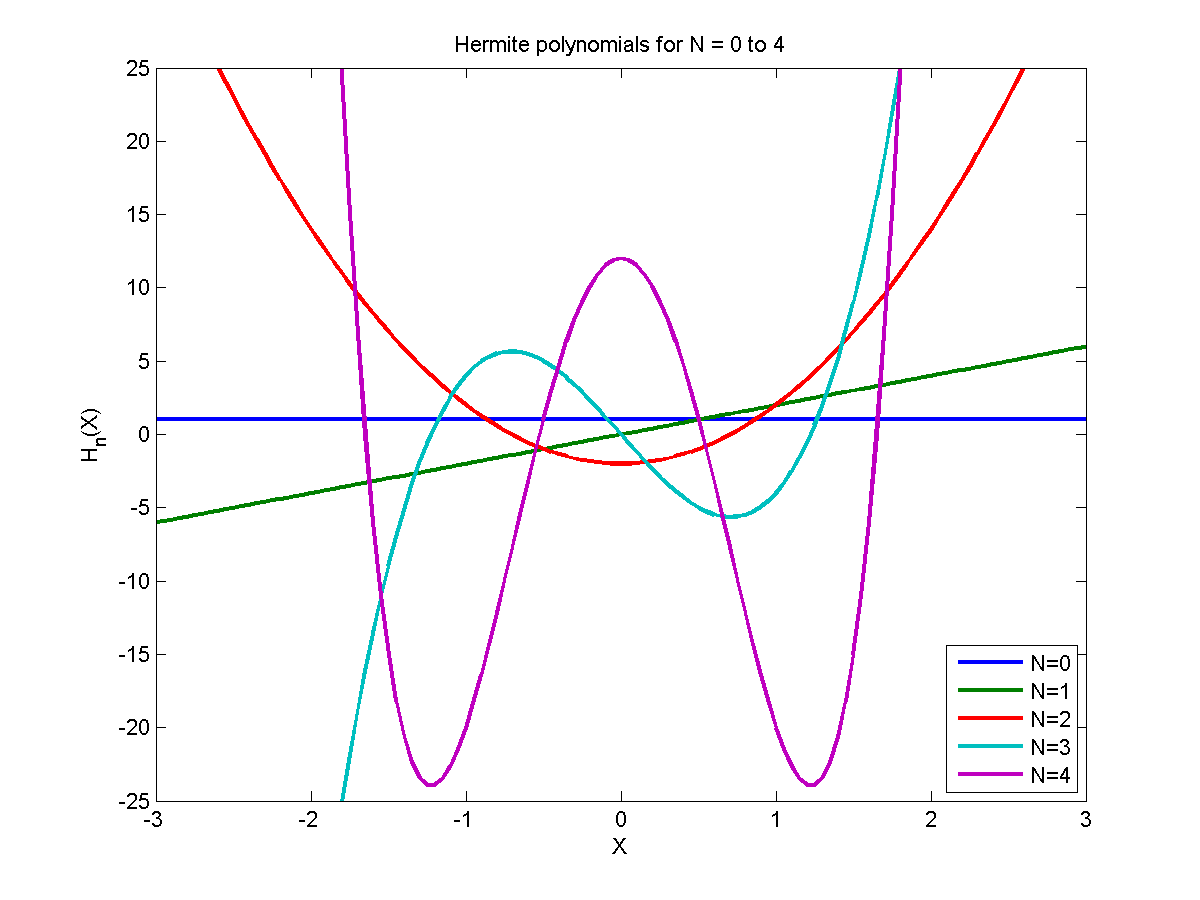
Orthogonal Polynomials
Standard polynomials can cause numerical issues due to differences in scale:
\(X = 100\) \(X^3 = 1,000,000\)
Centering and scaling \(X\) can help.
Alternatively, we can use “orthogonal polynomials” created using poly(raw=FALSE) (the default). See Section 4.10 in the book.
Splines
Species-Area relationship
Linear models are often a good approximation over small ranges of \(x\).
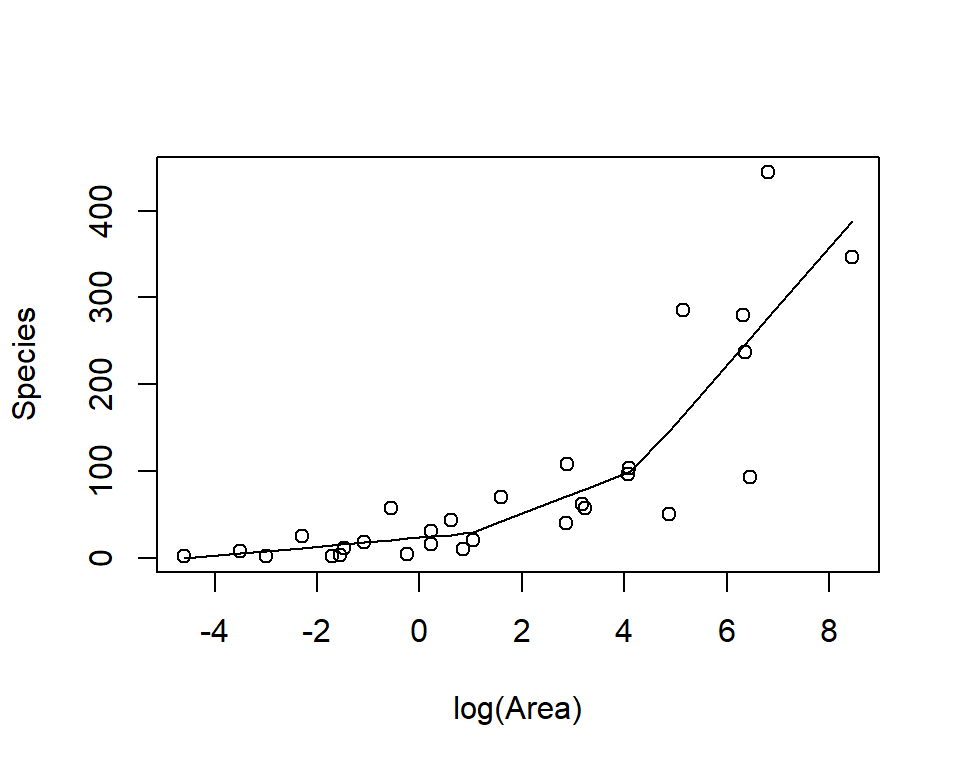
Splines
Splines are piecewise polynomials used in curve fitting.
A linear spline is a continuous function formed by connecting linear segments. The points where the segments connect are called the knots of the spline.
Linear spline with knots at 1 and 4.2
Call:
lm(formula = Species ~ logarea + logarea.1 + logarea.4.2, data = gala)
Residuals:
Min 1Q Median 3Q Max
-160.691 -16.547 -4.209 13.133 166.430
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 23.869 17.384 1.373 0.1819
logarea 5.213 8.956 0.582 0.5658
logarea.1 17.464 18.836 0.927 0.3627
logarea.4.2 44.815 23.156 1.935 0.0643 .
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 58.97 on 25 degrees of freedom
Multiple R-squared: 0.7695, Adjusted R-squared: 0.7418
F-statistic: 27.82 on 3 and 25 DF, p-value: 3.934e-08Basis functions
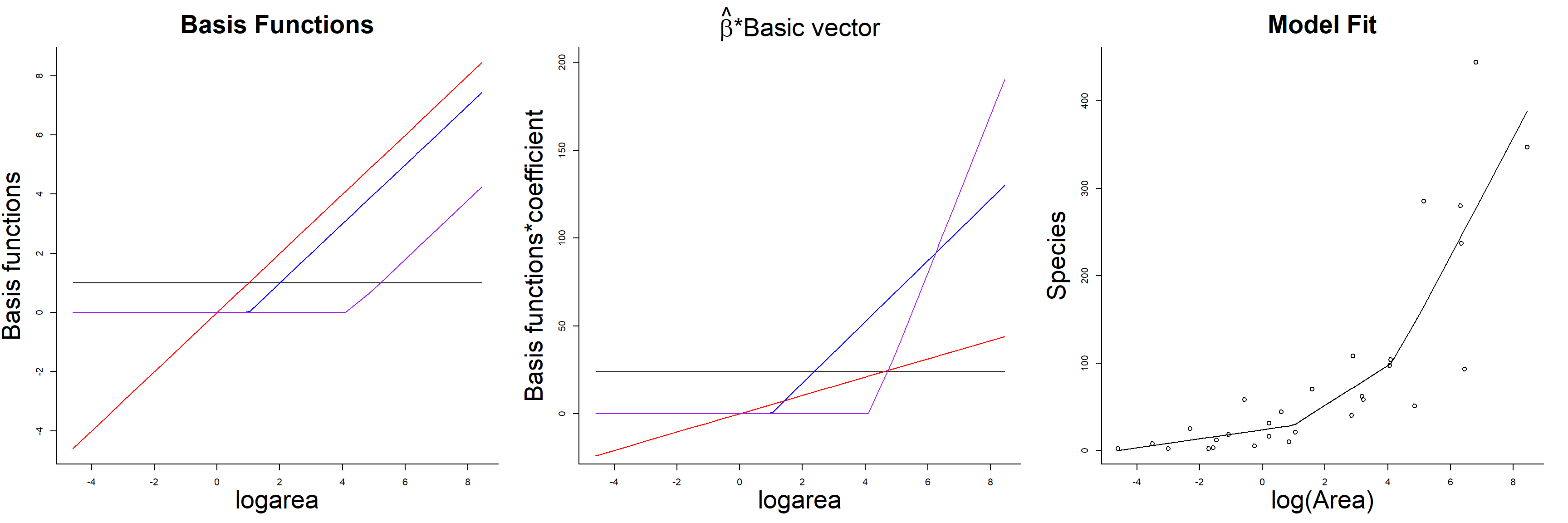
- Left = Basis Functions
- Middle = Basis Functions * regression coefficient
- Right = Fitted Model
Splines
A spline of degree D is a function formed by connecting polynomial segments of degree D so that:
- the function is continuous (no `jumps’)
- the function has D-1 continuous derivatives
- the D\(^{th}\) derivative is constant between knots
Linear splines (D = 1): first derivative is not constant (can go from increasing to decreasing at a knot)
Cubic Regression Splines
Fits a cubic polynomial on segments of the data
D-1 = 2 continuous derivatives everywhere (even at the knot locations)
- the first derivative (tells us if the function is increasing or decreasing) is continuous (even at the knots)
- the second derivative (tell us about curvature) is constant (even at the knots)
- Ensures that the fit is “smooth” at the connections (knot locations)
Truncated Power Basis
The truncated polynomial of degree D associated with a knot \(\xi_k\) is the function which is equal to 0 to the left of \(\xi_k\) and equal to \((x - \xi_k)^D\) to the right of \(\xi_k\).
\((x - \xi_k)^D_+ = \left\{ \begin{array}{ll} 0 & \mbox{if } x < \xi_k\\ (x - \xi_k)^D & \mbox{if } x \ge \xi_k \end{array} \right.\)
The equation for a spline of degree D with K knots is:
\(y = \beta_0 +\sum_{d=1}^D \beta_Dx^d + \sum_{k=1}^K b_k(x - \xi_k)^D_+\)
Splines
The design matrix for a cubic spline with K knots is the n by 1 + 3 + K matrix with entries:
\(\begin{bmatrix} 1 & x_1 & x_1^2 & x_1^3 & (x_1 - \xi_1)^3_+ & ... & (x_1 - \xi_k)^3_+\\ 1 & x_2 & x_2^2 & x_2^3 & (x_2 - \xi_1)^3_+ & ... & (x_2 - \xi_k)^3_+\\ 1 & x_3 & x_3^2 & x_3^3 & (x_3 - \xi_1)^3_+ & ... & (x_3 - \xi_k)^3_+\\ & & \vdots & & \\ 1 & x_n & x_n^2 & x_n^3 & (x_n - \xi_1)^3_+ & ... & (x_n - \xi_k)^3_+\\ \end{bmatrix}\)
Basis functions: Splines
Truncated power basis:
- Easiest to understand, but may run into numerical problems due to scaling issues
Bsplines ( in splines package)
- Numerically more stable than those based on the truncated power basis
- Can be poorly behaved in the tails
Natural or restricted cubic splines ( in splines package; in rcs package)
- Fit is constrained to be linear before the first knot and after the last knot (these are refered to as )
- Requires fewer model df (number of knots -1 = number of interior knots + 1)
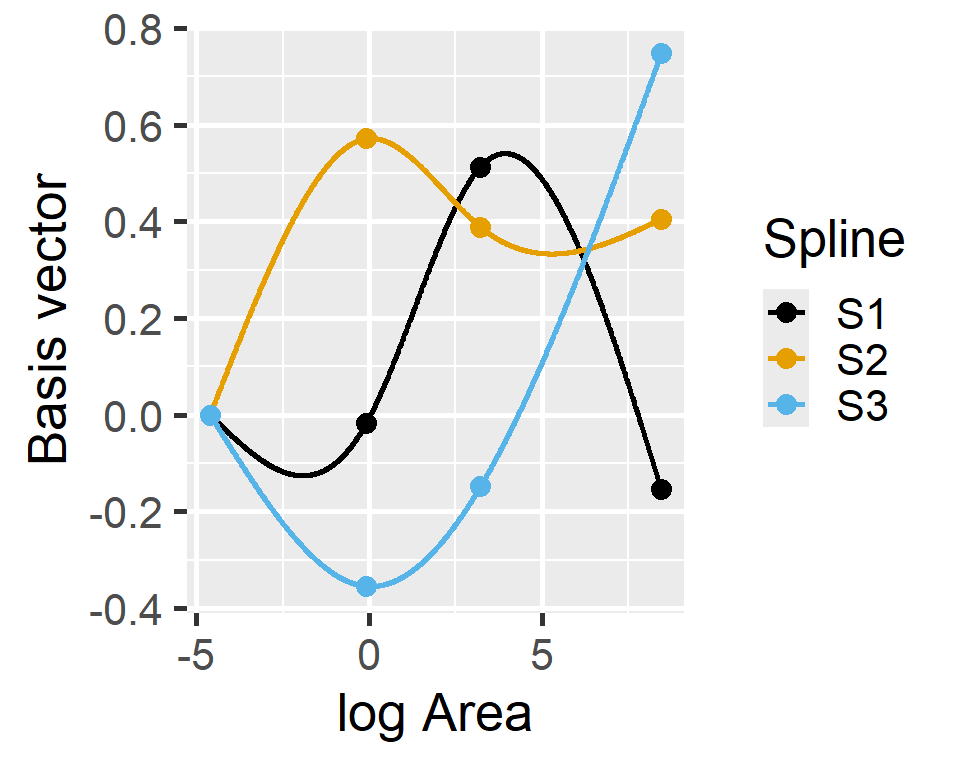
Natural Splines
Call:
lm(formula = Species ~ ns(logarea, df = 3), data = gala)
Residuals:
Min 1Q Median 3Q Max
-156.173 -13.819 -5.998 13.922 170.555
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 1.468 43.542 0.034 0.9734
ns(logarea, df = 3)1 47.790 45.957 1.040 0.3084
ns(logarea, df = 3)2 276.125 102.146 2.703 0.0122 *
ns(logarea, df = 3)3 381.743 45.084 8.467 8.22e-09 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 59.48 on 25 degrees of freedom
Multiple R-squared: 0.7655, Adjusted R-squared: 0.7374
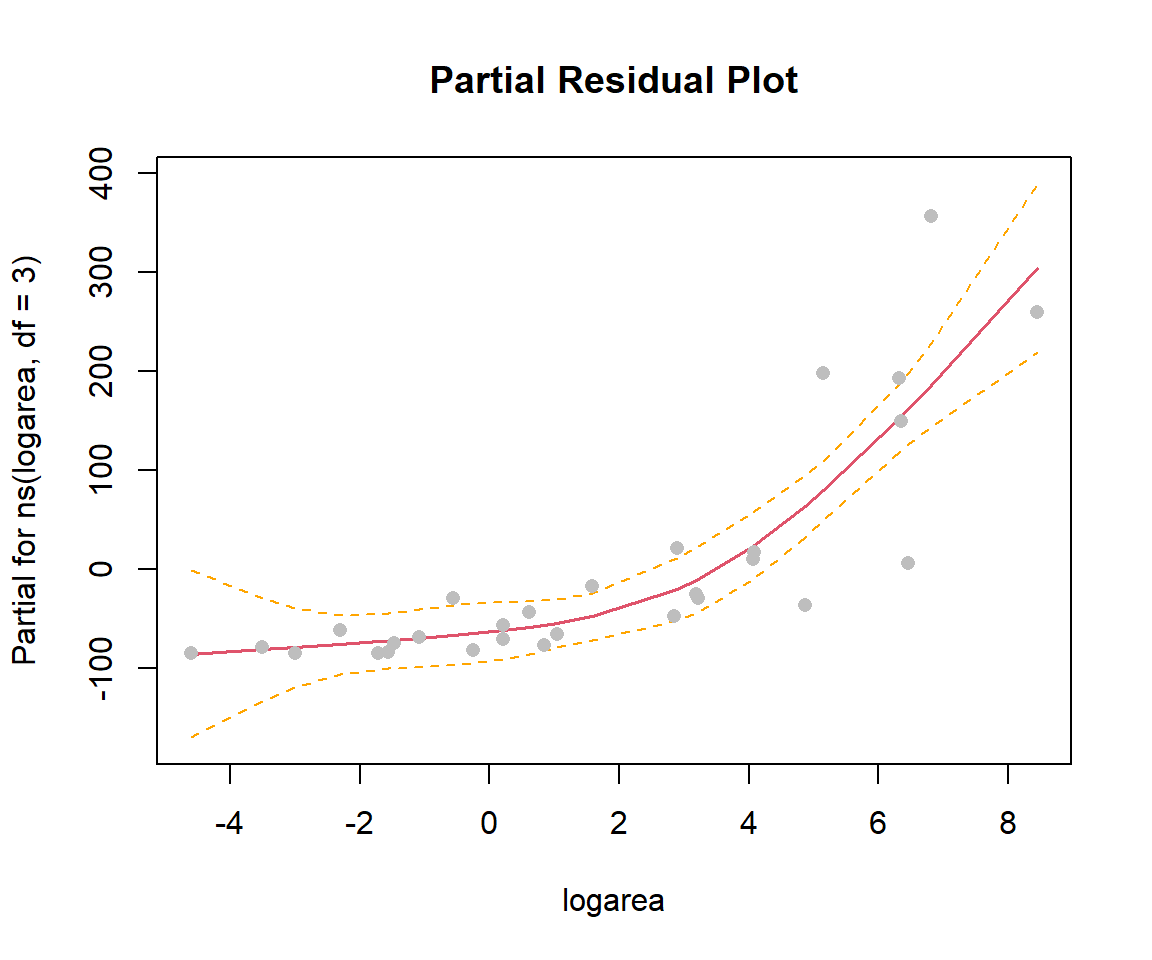
F-statistic: 27.21 on 3 and 25 DF, p-value: 4.859e-08Natural Splines: Basis Vectors
Natural Splines
Code
a <- ggplot(gala, aes(x = logarea, y = Species)) + geom_point(size = 3) +
geom_smooth(method = "lm",
formula = y ~ ns(x, 3),
se = TRUE) + ggtitle("Spline Model") + ylim(c(-100, 500))
b <- ggplot(gala, aes(x = logarea, y = Species)) + geom_point(size = 3) +
geom_smooth(method = "lm",
formula = y ~ poly(x, 2),
se = TRUE) + ggtitle("Polynomial Model")+ ylim(c(-100, 500))
gridExtra::grid.arrange(a, b, ncol = 2)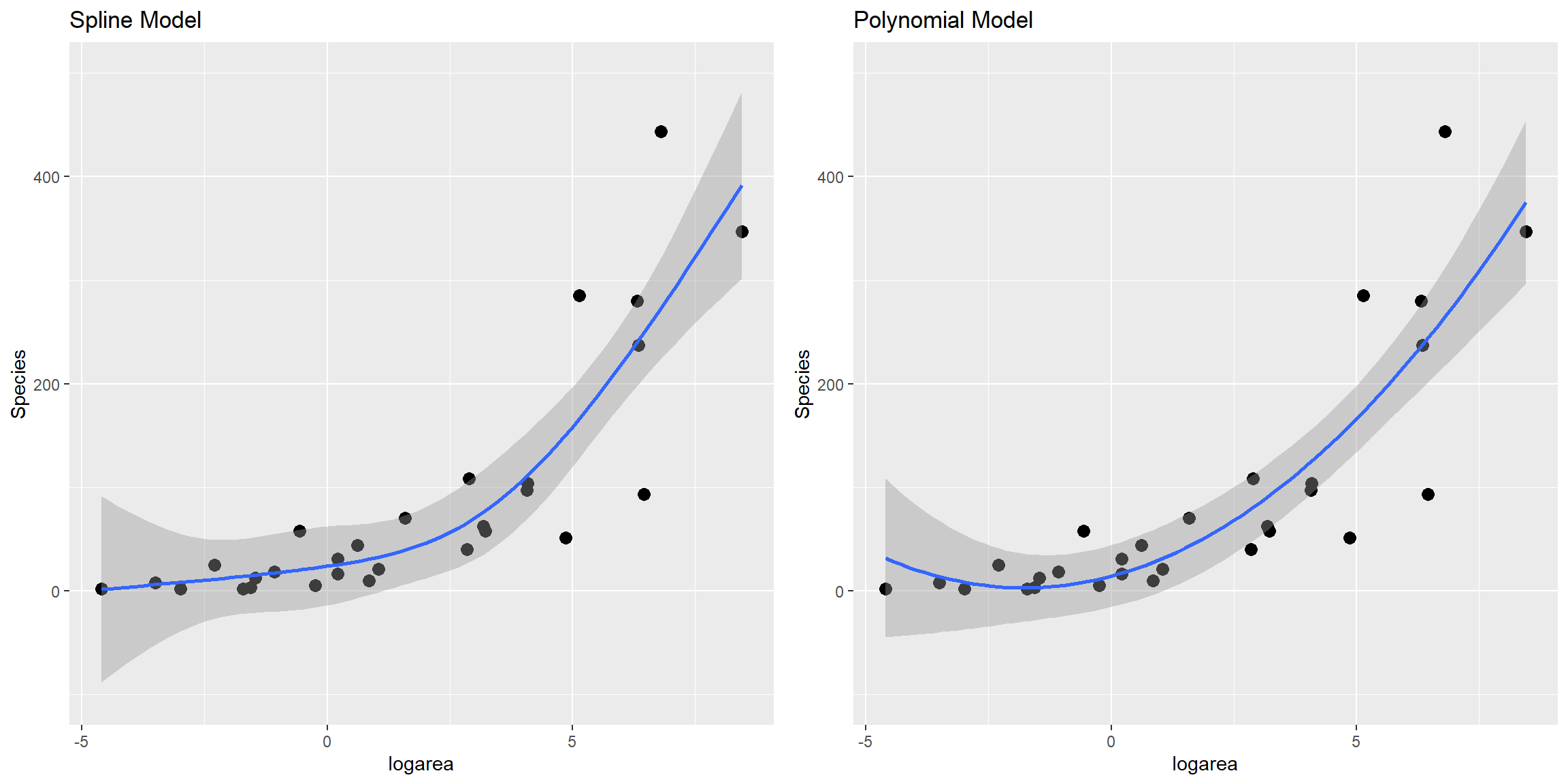
Natural Splines
Compare fit to that of linear model
df AIC
lmfit 3 335.1547
lm.poly1.raw 4 324.4895
lm.sp 5 324.4646
lm.ns 5 324.9600Any and all approaches fit better than a linear model!
Number of knots and their locations
The shape of a spline can be controlled by carefully choosing the number of knots and their exact locations in order to:
- Allow flexibility where the trend changes quickly, and
- Avoid overfitting where the trend changes little.
Could in principle compare models (e.g., using AIC) that have varying numbers of knots, or different knot locations
- Danger of overfitting, and difficult to account for model-selection uncertainty
Number of knots and their locations
Choose a small number of knots (df), based on how much data you have and how complex you expect the relationship to be a priori
- I’ve found that 2 or 3 internal knots are usually sufficient for small data sets
- Keele (2008), cited in Zuur et al, recommend 3 knots if \(n < 30\) and 5 knots if \(n > 100\)
Choose knot locations based on quantiles (what ns does by default if you do not provide knot locations)
- Models fit with cubic regression splines are usually not too sensitive to knot locations
Span: Splines
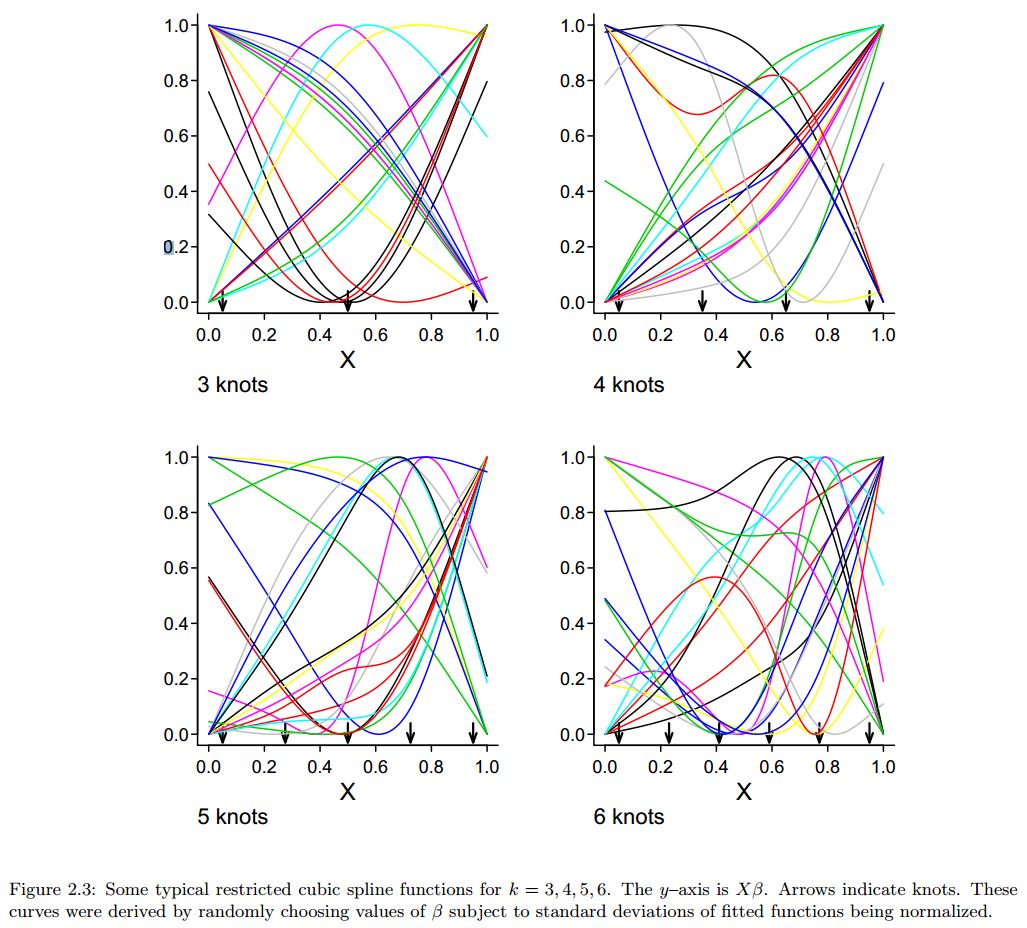
Generalized Additive Models
\(E[Y|X] = \beta_0 + f(x_1)\)
where \(f(x_1)\) can be modeled in a variety of ways
- Smoothing splines
- Loess (locally weighted linear regression)
Smoothing or Penalized Splines
Smoothing splines: Use lots of knots, but then attempt to balance overfitting and smoothness by controlling the size of the spline coefficients.
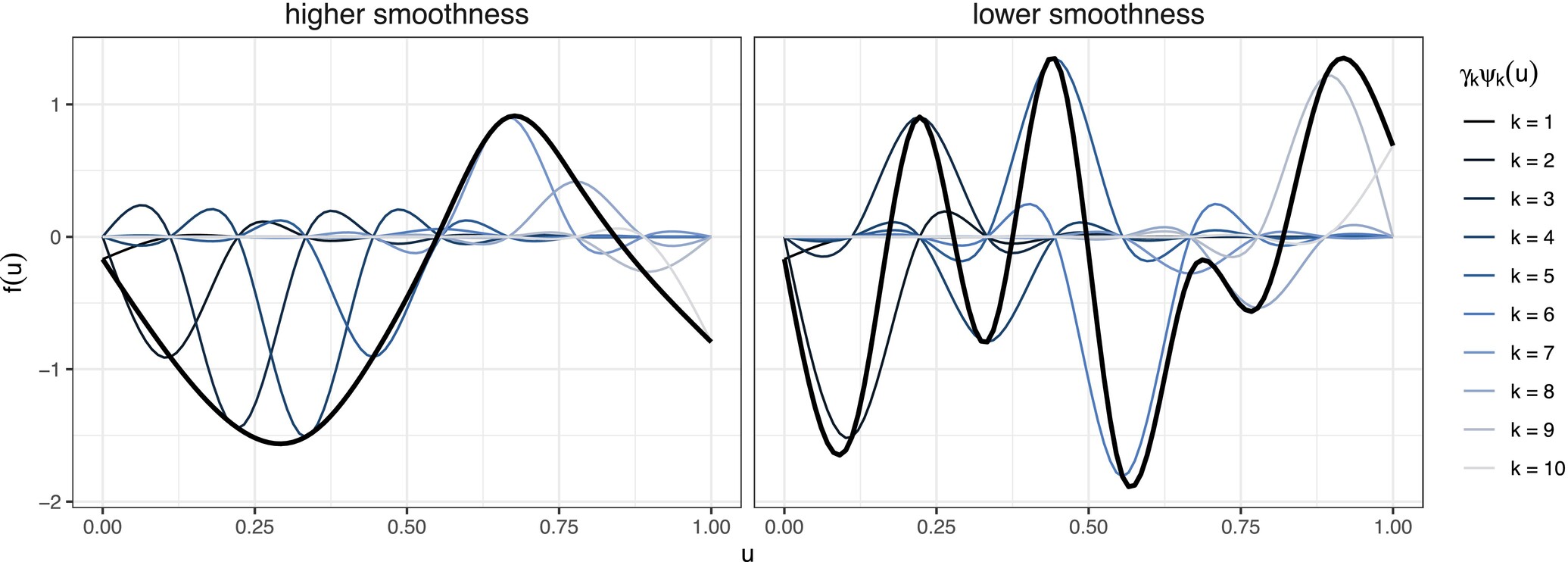
Klappstein, N. J., Michelot, T., Fieberg, J., Pedersen, E. J., & Mills Flemming, J. (2024). Step selection functions with non‐linear and random effects. Methods in ecology and evolution, 15(8), 1332-1346.
Other considerations
What if you want to allow for multiple non-linear relationships?
ns(x1, 3) + ns(x2, 4)or multiple smoothing splines
- Other basis functions can be used to fit “smooth surfaces” (allowing for interactions between variables)
- tensor splines, thin plate splines, etc…
- Can include interactions (separate smooth for each level of a categorical variable)
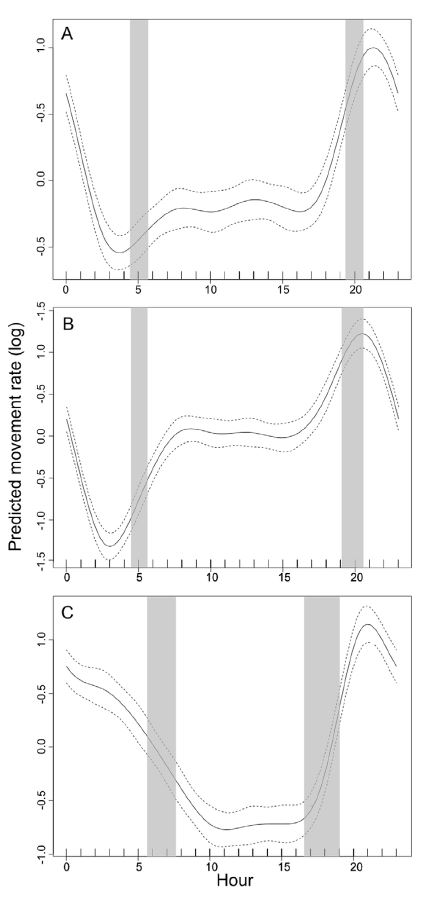
Black Bear Movement and Heart Rates

There are cyclical splines that
ensure ends meet at 0 and 24 hours
(or, Jan 1 and Dec 31).
Non-Linear Models with Mechanistic Basis
\(Y \sim f(x,\beta)\), where \(f(x,\beta)\) may have a strong theoretical motivation.
- Ricker model for stock-recruitment: \(S_{t+1} = S_te^{r(1-\beta S_t)}\)
- Predator prey: \(f(N) = \frac{aN}{1+ahN}\)
We will eventually learn how to fit these models using Maximum likelihood and Bayesian methods.