lm.ref <- lm(log(lj_length) ~ log(skull_length)*Species, data = fishmorph2)Notes on Mid-term
Multiple Regression problem
\[\log(\text{lower jaw length})_i \sim N(\mu_i, \sigma^2)\] \[\mu_i = \beta_0 + \beta_1\log(\text{Skull Length})_i + \beta_2I(Species=PL)_i + \beta_3I(Species=XA)_i +\] \[\beta_4\log(\text{Skull Length})_iI(Species=PL)_i+ \beta_5\log(\text{Skull Length})_iI(Species=XA)_i\]
- \(\beta_0\) = mean or expected log(lower jaw length) when log(skull length) = 0 and species is AP (Anoplarchus purpurescens)
- \(\beta_1\) = increase in the expected value of log lower jaw length as we increase log(skull length) by 1 when species is AP (Anoplarchus purpurescens)
- \(\beta_2\) = difference in intercept for Species = PL relative to the intercept for species = AP; \((\beta_0 + \beta_2)\) gives the mean or expected log(lower jaw length) when log(skull length) = 0 and species is PL
- \(\hat{\sigma}^2 = 0.0281^2\) tells us how much unexplained variability we have about the line.
Design matrix
For the final exam, make sure you can determine the design matrix this without the help of model.matrix.
(Intercept) log(skull_length) SpeciesPholis_laeta
1 1 2.0 0
2 1 1.5 0
SpeciesXiphister_atropurpureus log(skull_length):SpeciesPholis_laeta
1 0 0
2 1 0
log(skull_length):SpeciesXiphister_atropurpureus
1 0.0
2 1.5 (Intercept) SpeciesXiphister_atropurpureus log(skull_length)
1 1 0 2.0
2 1 1 1.5
SpeciesXiphister_atropurpureus:log(skull_length)
1 0.0
2 1.5Means coding
Call:
lm(formula = log(lj_length) ~ log(skull_length) * Species - log(skull_length) -
1, data = fishmorph2)
Residuals:
Min 1Q Median 3Q Max
-0.036489 -0.021110 -0.004249 0.022981 0.040137
Coefficients:
Estimate Std. Error t value
SpeciesAnoplarchus_purpurescens -1.32633 0.12749 -10.403
SpeciesPholis_laeta -1.57038 0.28260 -5.557
SpeciesXiphister_atropurpureus -0.94076 0.09015 -10.435
log(skull_length):SpeciesAnoplarchus_purpurescens 1.39942 0.05740 24.380
log(skull_length):SpeciesPholis_laeta 1.39505 0.13760 10.139
log(skull_length):SpeciesXiphister_atropurpureus 1.11998 0.03815 29.360
Pr(>|t|)
SpeciesAnoplarchus_purpurescens 2.96e-08 ***
SpeciesPholis_laeta 5.49e-05 ***
SpeciesXiphister_atropurpureus 2.84e-08 ***
log(skull_length):SpeciesAnoplarchus_purpurescens 1.76e-13 ***
log(skull_length):SpeciesPholis_laeta 4.17e-08 ***
log(skull_length):SpeciesXiphister_atropurpureus 1.14e-14 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.0281 on 15 degrees of freedom
Multiple R-squared: 0.9998, Adjusted R-squared: 0.9997
F-statistic: 1.273e+04 on 6 and 15 DF, p-value: < 2.2e-16Hypothesis tests
2.5 % 97.5 %
SpeciesAnoplarchus_purpurescens -1.598064 -1.0545893
SpeciesPholis_laeta -2.172736 -0.9680278
SpeciesXiphister_atropurpureus -1.132916 -0.7486064
log(skull_length):SpeciesAnoplarchus_purpurescens 1.277071 1.5217612
log(skull_length):SpeciesPholis_laeta 1.101771 1.6883305
log(skull_length):SpeciesXiphister_atropurpureus 1.038671 1.2012844If you used reference coding, then you need to use matrix multiplication (or a package like emmeans) to get the species-specific estimates and their uncertainty.
- need to account for the variance of each parameter
- and how parameters co-vary
See Section 3.12 in the book and also the GLS slides/application.
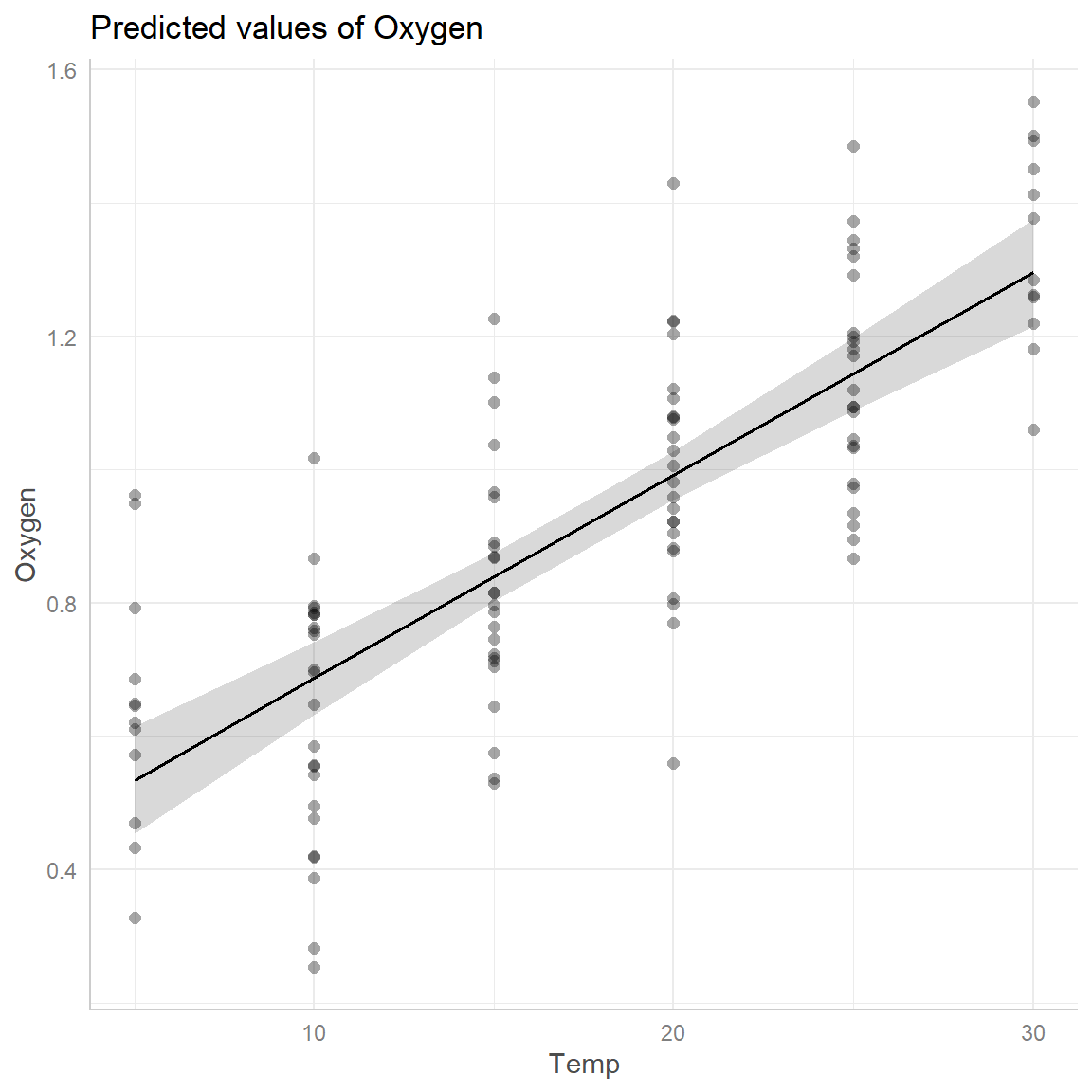
Non-linear regression
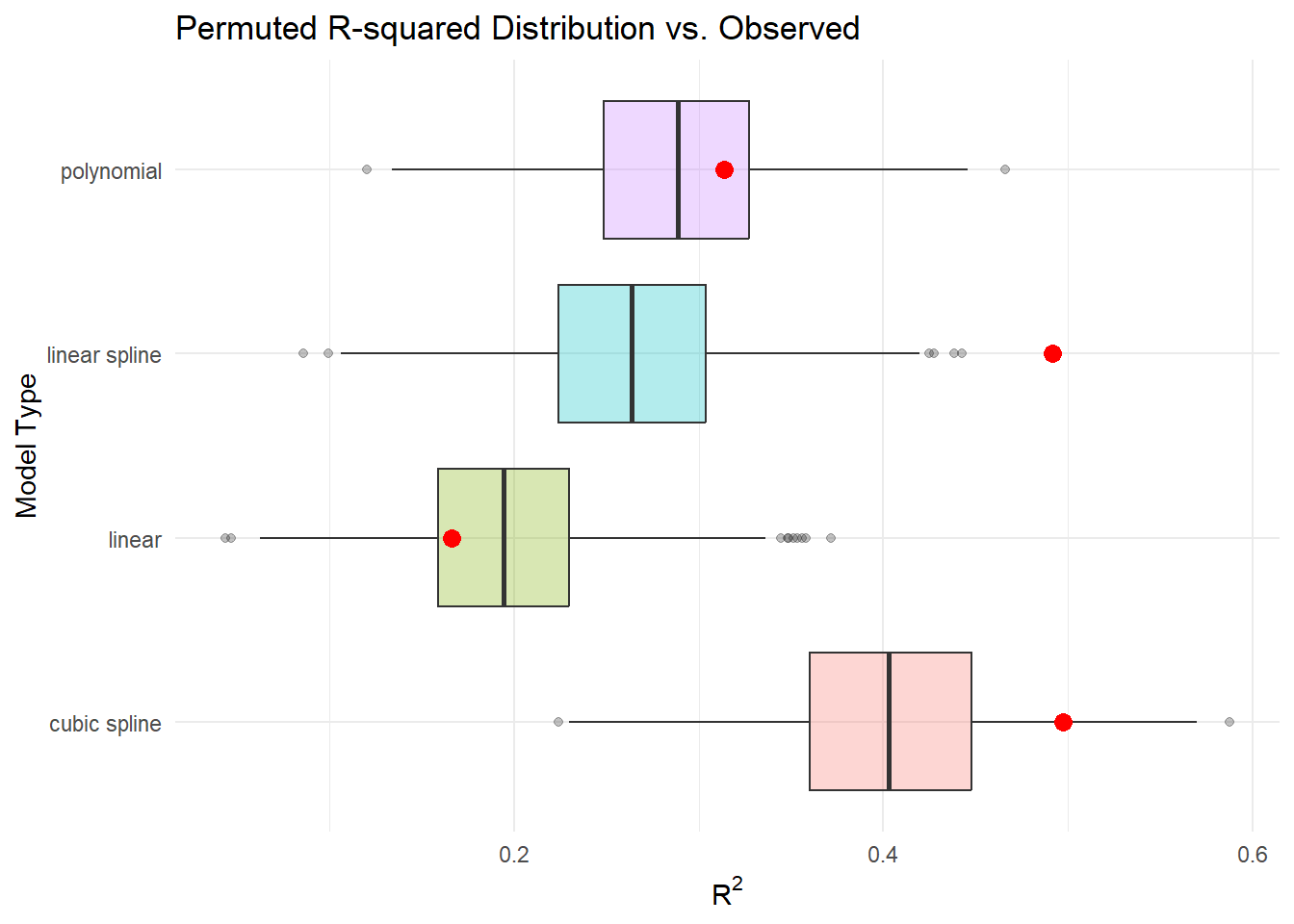
Out-of-sample \(R^2\)
We might expect out-of-sample \(R^2\) to be smaller than in-sample \(R^2\) if we use the data to inform our model in key ways:
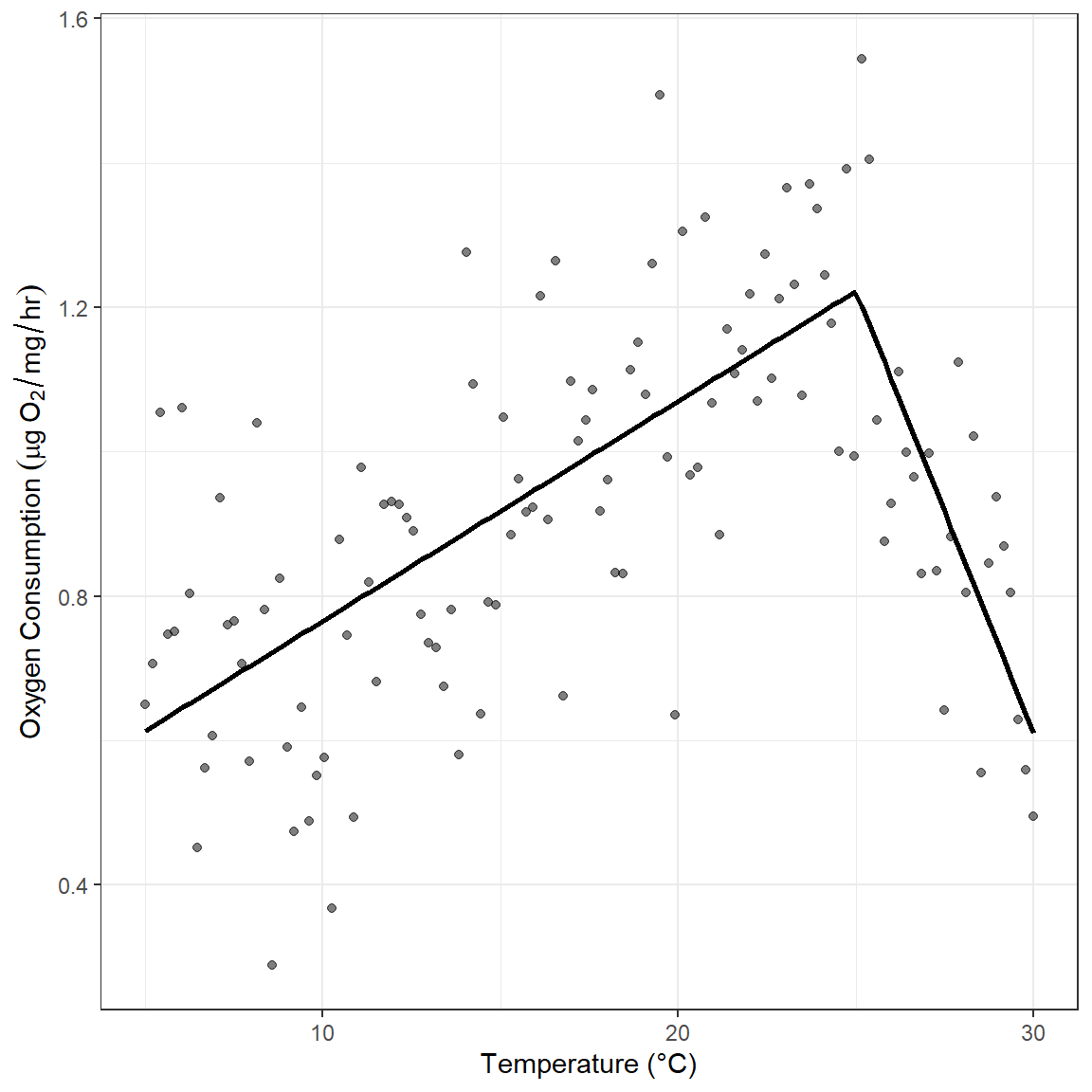
- pick the location of a knot for the linear spline
- determine how many df to allocate to cubic splines
- decide on a quadratic or cubic polynomial
This problem was not perfect, because ideally we would like to replicate the full process of:
- Collect a data set
- Use the data to inform our model
- Evaluate how the data-informed model performs on a new data set.
We didn’t replicate 1 and 2 (but replicated 3). But…
Results